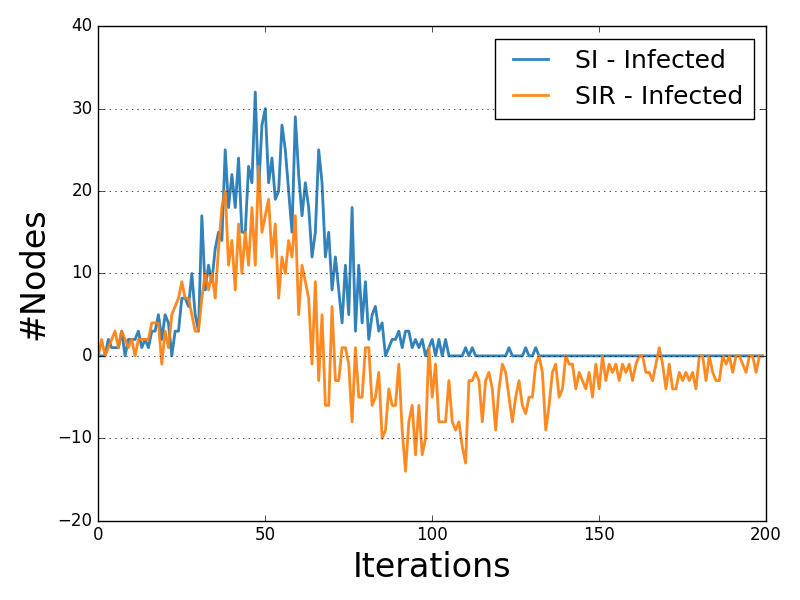
Diffusion Prevalence Comparison¶
The Diffusion Prevalence plot compares the delta-trends of all the statuses allowed by the diffusive model tested.
Each trend line describes the delta of the number of nodes for a given status iteration after iteration.
Below is shown an example of Diffusion Prevalence description and visualization for two instances of the SIR model.
import networkx as nx
import ndlib.models.ModelConfig as mc
import ndlib.models.epidemics as ep
from ndlib.viz.mpl.PrevalenceComparison import DiffusionPrevalenceComparison
# Network topology
g = nx.erdos_renyi_graph(1000, 0.1)
# Model selection
model = ep.SIRModel(g)
# Model Configuration
cfg = mc.Configuration()
cfg.add_model_parameter('beta', 0.001)
cfg.add_model_parameter('gamma', 0.02)
cfg.add_model_parameter("fraction_infected", 0.01)
model.set_initial_status(cfg)
# Simulation execution
iterations = model.iteration_bunch(200)
trends = model.build_trends(iterations)
# 2° Model selection
model1 = ep.SIModel(g)
# 2° Model Configuration
cfg = mc.Configuration()
cfg.add_model_parameter('beta', 0.001)
cfg.add_model_parameter("fraction_infected", 0.01)
model1.set_initial_status(cfg)
# 2° Simulation execution
iterations = model1.iteration_bunch(200)
trends1 = model1.build_trends(iterations)
# Visualization
viz = DiffusionPrevalenceComparison([model, model1], [trends, trends1])
viz.plot("trend_comparison.pdf")